
Current and evolving biomarkers for precision oncology in the management of metastatic colorectal cancer
Introduction
As the 3rd most common cancer worldwide, colorectal cancer (CRC) is the 2nd leading cause of death, accounting for an estimated 1.8 million new diagnoses and 880,000 deaths globally in 2018. Worldwide, the incidence of CRC diagnoses varies significantly with rates almost 4-fold higher in regions with high human development index (HDI). Three distinct global trends have been observed and linked to levels of regional development with increases in incidence and mortality in Russia, China and Brazil, with increasing incidence but decreasing mortality in the UK and Singapore, and with decreasing incidence and mortality in the United States and Japan (1,2). Rises in incidence have been attributed to worldwide dietary changes, obesity and lifestyle factors. Conversely, declining mortality has been linked to robust screening patterns and adoption of best practices (3-5). While in the United States CRC rates have been decreasing by about 2 percent per year population wide, the incidence of patients with CRC younger than age 50 is increasing. Colon cancer and rectal cancer are estimated to increase by 90 percent and 124 percent, respectively in patients 20–34 years old by 2030 (6,7).
Approximately 50–60% of patients who are diagnosed with CRC will eventually develop metastatic disease (8). Most often, metastases develop after treatment for locoregional disease, however 20–34% of those with CRC will present with synchronous metastatic disease. Unfortunately, the majority of patients who present with metastatic CRC (mCRC) have inoperable disease (9-11). The standard of care for these patients is antineoplastic agents with an ultimate goal of improving quality of life and prolonging survival.
5-fluorouracil represented the mainstay of treatment in patients with mCRC from 1962 until 1996. In the span of the following 6 years, the FDA approved three new cytotoxic agents starting with irinotecan, followed by capecitabine in 2001 and oxaliplatin in 2002 (12). Since 2002, there have been 13 new FDA drug approvals and 5 additional drugs that appear on the National Comprehensive Cancer Network (NCCN) guidelines but still await FDA approval. Therefore, the current arsenal of drugs include monoclonal antibodies (mAbs) that target anti-epidermal growth factor receptor (EGFR) (cetuximab and panitumumab), vascular endothelial growth factor/vascular endothelial growth factor receptor (VEGF/VEGFR) (bevacizumab, ramucirumab, and ziv-aflibercept), small molecule tyrosine kinase inhibitors (TKIs) (regorafenib, binimetinib/encorafenib, trametinib/dabrafenib, vemurafenib, larotrectinib), and immune checkpoint inhibitors, including mAbs targeting anti-programmed death-1 (PD-1, nivolumab and pembrolizumab) as well as anti-cytotoxic T-lymphocyte antigen-4 (CTLA-4, ipilimumab). Trifluridine/tipiracil is the only cytotoxic agent that has been approved since 2002 (10).
With significant advances in systemic chemotherapeutic and biologic options for mCRC, outcomes have improved with median overall survival (OS) now exceeding 30 months (10,11). However, long term outcomes still remain poor with 5-year OS for patients with metastatic disease estimated at only 14.2%, declining to 7.4% in patients older than 65 (13,14). While continual clinical investigation for novel treatment modalities remains imperative, honing in on distinct molecular subsets of CRC has allowed us to leverage targeted therapy in a meaningful way. Over the last several years, more robust research has demonstrated that mCRC is not one entity, but rather consistent with significant molecular heterogeneity (Figure 1). These advances in understanding the molecular and cellular mechanisms of mCRC increasingly affect prognosis and treatment decisions. Furthermore, this underlying biologic heterogeneity explains the tremendous variability noted in regards to treatment outcomes, especially in the advanced setting. Today, molecular biomarkers are a critical component of the management of mCRC, serving as both prognostic and predictive tools. In light of this, we are becoming progressively aware of the proper utilization of pharmacologic agents at our disposal. In this review, we will summarize the current literature on molecular biomarkers in CRC and their impact on treatment decision making, as well as highlight evolving biomarkers of increasing clinical significance.
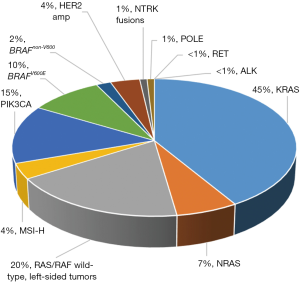
Established molecular biomarkers in CRC
RAS
The mitogen-activated protein kinase (MAPK) pathway, altered in many cancers, is essential in a number of cellular pathways. Dysregulation leads to uncontrolled cellular proliferation, survival and dedifferentiation. The first step in initiation of the MAPK pathway occurs when ligands such as growth factors, cytokines or hormones bind to extracellular membrane receptor tyrosine kinases such as EGFR. Binding activates RAS, which activates BRAF, followed by MEK kinase (MEK1 and MEK2), and finally ERK (15).
The rat sarcoma viral oncogene (RAS) was discovered in 1982 and marked the first discovered mutated gene in disease (16). Since that time, there have been over 500 validated genes in cancer (17), but the three RAS genes (HRAS, NRAS, and KRAS) still represent the most mutated oncogene family in cancer (30%) and an estimated 52% of all CRCs. KRAS is the most frequently isolated form, representing 86% of RAS mutations in CRC, followed by 14% NRAS mutations (18). The majority of KRAS mutations in colon cancer affect codons 12 (30%) and 13 of exon 2 (8%). An additional 6% of mutations are found in KRAS exons 3 and 4, while 5% of mutations are in NRAS exons 2, 3 or 4 (19).
RAS genes encode a 21-kDa monomeric GTPase downstream of EGFR that transduces extracellular signal to intracellular signals by binding growth factors to cell membrane receptors and helps mediate signals related to cell survival, senescence and other cell survival signals.
They are essential components of the EGFR signaling cascade, and mutations can isolate the pathway from EGFR signaling. RAS proteins are GTP-binding proteins, with GTPase functionality that switches between the active and inactive state. Missense mutations in RAS proteins alter binding toward the active state (20). While complex, the two major signaling pathways that RAS proteins affect are the MAPK and the phosphoinositide-3 kinase (PI3K) pathways.
Increasing understanding of CRC carcinogenesis has led to the production of mAbs that target EGFR. EGFR is a commonly expressed transmembrane glycoprotein that is a member of the human epidermal growth factor receptor (HER) tyrosine kinase growth factor receptor family and is encoded by proto-oncogene c-erb B. Activation of c-erb B proto-oncogene results in expression of EGFR, which prompted interest as a potential target for anticancer therapy (21). Upon extracellular ligand binding to growth factors, receptor dimerization occurs, and the intracellular tyrosine kinase is activated, prompting downstream signaling, including RAS (22,23).
EGFR is often upregulated in CRC and presents an opportunity for therapeutic intervention by preventing signal transduction of dependent pathways involving RAS, PI3K and MAPK pathway. With this strategy in mind, two mAB’s, cetuximab and panitumumab, were tested and found to improve OS in patients with metastatic colon cancer. Cetuximab (a chimeric IgG1 mAb) and panitumumab (a fully human mAb) both bind to the extracellular domain of EGFR, and block ligand induced receptor signaling and therefore tumor growth (24,25). Shortly after drug approval, there were a number of early studies that suggested that mutations in KRAS conferred resistance to cetuximab (26,27). Post hoc analysis of both CRYSTAL (FOLIRI +/− cetuximab) and PRIME (FOLFOX +/− panitumumab) demonstrated irrevocable resistance when RAS-mutant mCRC patients receive anti-EGFR mAb therapy (27,28). As we have learned, and discuss further below, only 10–20% of patients can expect benefit from EGFR inhibition (EGFRi) (10).
RAS wild type (WT) and tumor sidedness
Primary tumor location in mCRC has emerged as a prognostic and potentially predictive tool. The NCCN defines right sided tumors as those that arise in the cecum to the hepatic flexure, while left sided tumors are those from the splenic flexure to rectum. Initial signals reflecting the impact of sidedness were noted in a first-line chemotherapy study revealing right sided tumors exposed to 5-FU based chemotherapy had worse outcomes than left sided tumors, with a difference of at least 5 months in survival (29). These findings have since been reproduced in a more recent analysis of two randomized phase III trials, AVF2107g and NO16966. Both trials utilized frontline chemotherapy in combination with bevacizumab for treatment-naïve mCRC. Tumor location analysis revealed right sided tumors as a negative prognostic variable irrespective of mucinous histology or mutational status, with agnostic efficacy of bevacizumab (30). A post hoc analysis of RAS WT populations in the CRYSTAL, PRIME, PEAK and FIRE-3 trials found that the benefit of anti-EGFR therapy in terms of OS and progression-free survival (PFS) remained only for patients with left sided tumors (31-34). The most convincing evidence for the predictive lack of response of EGFR inhibitors in regards to tumor sidedness stems from the CALGB/SWOG 80405 study. The authors found that among RAS WT patients treated with chemotherapy + cetuximab, there was a statistically significant reduction in OS of 13.6 months for right sided tumors in contrast to 39.3 months for left sided tumors [P=0.001, hazard ratio (HR) 0.55] (35). Bevacizumab exposure, however, remained agnostic in terms of survival outcomes for RAS WT left vs. right sided tumors (P=0.50).
Current guidelines, both from the European Society of Medical Oncology (ESMO) and NCCN, recommend against treating patients with RAS-mutant mCRC with anti-EGFR therapy, either alone or in combination with chemotherapy. In regards to RAS-WT tumors, due to the retrospective findings of CALGB 80405 highlighting sidedness as a reliable biomarker of response and selection for anti-EGFR therapy, a paradigm shift in the management of mCRC has emerged. In light of this practice changing data, the NCCN definitively supports excluding patients with RAS-WT right sided tumors from exposure to EGFR inhibitors in the first-line setting (ESMO and NCCN). However, whether or not right sided RAS-WT patients should be treated with EGFRi in subsequent lines of therapy, remains an area with limited data and therefore a lack of consensus guidelines. Nonetheless, in the landmark CO.17 trial that confirmed lack of benefit of cetuximab compared to best supportive care for refractory KRAS mutant tumors, there was no statistically significant benefit with cetuximab among RAS WT right sided colon cancers compared to left sided colon cancers (36). Additionally, two smaller studies showed a similar trend for lack of benefit with anti-EGFR therapy in second line and beyond for right sided tumors; however, these cohorts are too small for definitive conclusions (37). Further studies are required to elucidate whether anti-EGFR therapy is appropriate in subsequent lines of therapy for patients with RAS WT right sided CRC.
BRAFV600E
BRAF is a serine threonine kinase downstream of RAS in the MAPK pathway and almost always exclusive of RAS mutations (10). BRAF mutations in CRC consists of a valine to glutamic acid change at codon 600 (c.1799T>A or p.V600E), and mutations result in a constitutively active protein, representing 96% of all BRAF V600 mutations (17,38). While BRAFV600E mutations are relatively uncommon in CRC with an estimated incidence of 7–14%, their extremely poor prognosis has warranted intense clinical and research focus (10,11,39,40). Of note, BRAFV600E mutations have distinct clinicopathologic factors including right sided tumors, high grade, older age, female sex, T4 tumors, mucinous histology, poorly differentiated tumors, and microsatellite instability (41-43). They tend to have higher rates of peritoneal disease, distant nodal involvement and brain metastases, but a lower incidence of lung metastases (44). Tumorigenesis in CRC occurs via specific molecular pathways, most commonly the classical (APC, KRAS, p53) pathway or the germline mutation (Lynch syndrome) pathway. However, the BRAF mutation is associated with a serrated adenoma precursor via the serrated/methylated pathway manifesting in a hypermethylated phenotype, which can result in inactivation of MLH1 causing a mismatch repair (MMR) deficiency and microsatellite instability (40,45). BRAF mutational status remains a strong prognostic factor for OS in both the metastatic setting and early stage disease (46). In the adjuvant setting, patients with BRAFV600E CRC tend to have shorter disease-free survival and worse OS after recurrence (47). Unsurprisingly, these patients tend to have diminished response to therapy with a median OS of around 12 months (48-50).
While the negative prognostic role of BRAFV600E is clear, historically, its predictive role in the first-line setting is less established. Initial retrospective studies were conflicting in terms of benefit to anti-EGFR therapy (51,52). For example, an unplanned retrospective subset analysis demonstrated that patients whose tumors were treated with FOLFIRI + cetuximab vs. FOLFIRI alone who were KRAS WT/BRAF mutant, had an improved OS of 14.1 vs. 10 months (53). Additionally, a planned subset analysis of the PRIME study demonstrated that BRAFV600E mutations were not predictive of benefit to panitumumab added to FOLFOX (54). Conversely results from the COIN trial suggested no benefit with addition of cetuximab to FOLFOX with a trend towards harm (55). In subsequent lines of therapy, the lack of benefit from mABs is clearer. One retrospective study demonstrated that patients with BRAFV600E had only an 8.3% response rate to cetuximab (56). Additionally, data from the PICCOLO study suggested harm with the addition of panitumumab to irinotecan in subsequent line settings (57). Of note, a large meta-analysis of 9 phase III and 1 phase II trial (six 1st line trial, two 2nd line trials, two chemotherapy refractory trials) evaluated a total of 463 pooled BRAFV600E patients. This analysis revealed that the addition of anti-EGFR therapy to standard chemotherapy or best supportive care did not significantly improve PFS, OS, or overall response rate in patients with BRAFV600E mCRC compared to control regimens. Although this analysis remains limited by size, being underpowered, non-randomized and retrospective in nature, there remains a notable signal that exposure to anti-EGFR therapy has limited clinical efficacy both as monotherapy or in combination with chemotherapy in BRAFV600E mCRC (58). In line with this data, both the NCCN and ESMO suggest against the utilization of EGFRi in isolation or in combination with chemotherapy for patients with BRAFV600E mCRC (10,11).
Unlike RAS/RAF WT mCRC, patients who harbor BRAFV600E mutation do not benefit from exposure to multiple lines of chemotherapy, with historical median PFS of 2.5 months among those who received second- and third-line treatment (59). Therefore, initial therapeutic decisions for this unique patient population remain critical. There is a growing body of evidence demonstrating viable alternate treatment approaches for these patients. The TRIBE study compared bevacizumab + FOLFIRI to bevacizumab + 5-FU + oxaliplatin + irinotecan (FOLFOXIRI) for treatment naïve unresectable mCRC. The study met its primary endpoint of improved PFS with an increase by 2.4 months for patients randomized to the intensified triplet arm compared to standard doublet chemotherapy. Of note, there were 28 patients in this study with BRAFV600E mutant disease of whom 16 received FOLFOXIRI + bevacizumab (B). While numbers are small in this subgroup analysis, patients in the FOLFOXIRI + B arm had an improved OS of 19 vs. 10.7 months in the FOLFIRI + B arm, consistent with historical norms (60). Based on these results, FOLFOXIRI + B remains the best chemotherapeutic option for patients with BRAFV600E mutation and adequate performance status in the first-line setting. It is sobering to note that comparatively RAS/RAF WT patients exposed to triplet chemotherapy had a median OS of 41.7 months in this study, highlighting the need for novel therapeutic strategies for this chemotherapy refractory subtype.
With encouraging results in melanoma, interest increased in BRAF inhibition in mCRC. Unfortunately, early studies demonstrated very little activity of vemurafenib and encorafenib with response rates of 5% and 0% respectively (61,62). This finding was somewhat unsurprising as there was little benefit in pre-clinical studies. However, it was discovered that while BRAF inhibition transiently induces impairment in the MAPK pathway, rapid reactivation of ERK occurs through activation of RAS and RAF (40). With this mechanism in mind, early phase trials demonstrated modest but improved clinical efficacy with combined BRAF and MEK inhibition with dabrafenib and trametinib and a response rate of 12%, including one complete response (63). Expanding on the concept of dual inhibition, the Southwest Oncology Group (SWOG) 1406 randomized 106 previously treated mCRC patients to irinotecan, and cetuximab, with or without vemurafenib, allowing crossover at progression. The authors found an improved PFS of 4.4 vs. 2.0 months and response rates of 16% vs. 4% in the vemurafenib arm (64).
More recently the phase III BEACON study, a 3-arm trial for treatment refractory BRAFV600E mCRC comparing a BRAF inhibitor (encorafenib) + MEK inhibitor (binimetinib) + cetuximab vs. encorafenib + cetuximab vs. standard of care (irinotecan/FOLFIRI +cetuximab), demonstrated impressive improvement on previous strategies by meeting primary endpoints of ORR and OS. Of note, based on a press release on 5/21/19, the OS HR was reported at 0.52 with improved response rate of 26% (vs. 2% in control) and an even higher response rate noted in 2nd line (34%) (65). Based on these exciting results, the NCCN has recommended the BEACON regimen of encorafenib in combination with binimetinib and cetuximab and triplet regimens of dabrafenib/trametinib/EGFR mAb as available treatment options for BRAFV600E mCRC, representing for the first time a completely targeted therapeutic approach for these high-risk patients (10).
Microsatellite-instability high (MSI-H)
MSI-H or deficient MMR (dMMR) tumors represent approximately 4–5% of all mCRC patients with a vastly different prognosis and exciting additional treatment options in the form of immunotherapy (10,11,66,67). Microsatellites are repetitive DNA sequences which represent a large portion of our genome and are susceptible to errors in insertion and deletions during DNA replication. During DNA synthesis, the primer and template strands in a microsatellite may become mismatched and the number of repeating strands may differ between the two strands. In normal functioning cells, these errors are reconciled by the MMR system. When MMR is defective, mistakes are replicated resulting in MSI and hypermutations (68).
Mutations in MMR may be hereditary or acquired. Germline mutations in MMR genes including MLH-1, MSH2, MSH6 and/or PMS2 or EpCAM are mutations found in patients with hereditary nonpolyposis CRC, also known as Lynch syndrome, and represent 1–6% of all CRCs (69). Somatic MMR defects are estimated to occur in 19% of all CRC patients and are associated with higher rates of tumor infiltrating lymphocytes (70). As mentioned above, the rates of patients with MSI-H and metastatic disease are much lower with studies ranging from 3.5–5.5% of patients (71,72). Intriguingly, multiple studies suggest improved prognosis for patients with stage II CRC. Specifically, the QUASAR study demonstrated a reduced risk of recurrence of 11% for those with dMMR in contrast to 26% for those with proficient mismatch repair (pMMR) (73).
Tumors with dMMR contain thousands of mutations which allow for easier recognition and targeting by the immune system. With the advent of checkpoint inhibitors, this has made for an intriguing new therapeutic strategy for treating this subset of patients. Specifically, strategies targeting programmed death ligand 1 (PD-L1) on tumor cells or its counterpart PD-1 on T cells have been evolving. Pembrolizumab and nivolumab, two IgG4 mAbs targeting PD-1, have shown promising, practice changing durable activity. Nivolumab was investigated in the Checkmate-142 in two separate cohorts, as single agent or in combination with ipilimumab, a fully human IgG1 mAb that targets the CTLA-4. In the single agent arm, including heavily pre-treated patients, the objective response rate (ORR) was 31.1% with 69% of patients demonstrating disease control at 12 weeks, with a median OS of 73% at 1 year (74). In the combination cohort, the ORR was 55% with a disease-control rate of 80% and an OS of 85% at 1 year. The combined efficacy came at a cost of 32% grade 3–4 treatment related events (75). Pembrolizumab also has demonstrated activity as a single agent in a recent phase II study. The study investigated pembrolizumab in tumors with MMR deficiency and included 11 patients with dMMR mCRC, 21 patients with pMMR CRC and an additional 9 patients with dMMR in the CRC. In the patients with mCRC dMMR group, a response rate of 40% and a PFS of 78% at 20 weeks were observed (67). This data, in conjunction with the nivolumab data, is indicative that MSI is a predictive marker for response to checkpoint blockade.
With this data in mind, both the NCCN and ESMO guidelines suggest universal testing for MMR or MSI with a dual purpose of identifying patients with Lynch syndrome to provide prognostic information for patients with stage II disease and to advise on use of immunotherapy in the advanced setting. NCCN recommends pembrolizumab, nivolumab or nivolumab plus ipilimumab as recommended treatment options for patients with advanced or metastatic dMMR CRC in second or third-line settings or for those patients “not appropriate for intensive therapy” (10,11).
HER2 amplified
HER2 is a member of the EGFR receptor family, and altered signaling may be caused by genomic amplification of ERBB2 or mutations. Activation leads to upregulation of the MAPK and PI3K pathways (76). HER2 amplifications are seen in 3–5% of all CRC, are mutually exclusive from RAS/RAF mutations, and have been associated with resistance to EGFR inhibitors (77). ERBB2 is far more commonly amplified in breast cancer and has been exploited as a target with therapeutic efficacy with mAbs such as trastuzumab and pertuzumab. Similar approaches are emerging in CRC and potentially represent a new therapeutic approach for these patients. The HERACLES study enrolled patients with KRAS WT, and HER2 positivity defined as tumors with 3+ HER2 score in more than 50% of cells by immunohistochemistry or with 2+ HER2 score and a HER2:CEP17 ratio higher than two in more than 50% of cells by fluorescence in situ hybridization (FISH). Patients were treated with trastuzumab (HER2 mAb) and lapatinib (an oral dual HER2/EGFR kinase inhibitor) after failure of standard therapies. Twenty-seven patients were enrolled and 8 patients (30%) had an objective response with one patient obtaining a complete response (4%) (78). More recently, the phase IIa MyPathway, a multiple basket study, reported their data on treatment-refractory, histologically confirmed HER2-amplified mCRC treated with dual HER2-targeted therapy with pertuzumab plus trastuzumab. Fifty-seven eligible patients were enrolled with 18 patients having an objective response (32%), one of which obtained a complete response (79). Based on these results the NCCN has now recommended the use of trastuzumab and lapatinib or trastuzumab and pertuzumab for HER2-amplified refractory mCRC patients (10). Considering the data revealing HER2 amplification as a negative predictive biomarker for anti-EGFR therapy, the SWOG1613 (NCT03365882) is an ongoing phase II clinical trial for anti-EGFR naïve RAS/RAF WT mCRC patients who have received at least 2 prior lines of therapy, randomizing patients to trastuzumab and pertuzumab vs. cetuximab and irinotecan.
NTRK fusions
Neurotrophic receptor tyrosine kinase fusion genes (NTRK) encode for TrkA, TrkB, and TrkC receptor tyrosine kinases that are important in the function of the nervous system in human neuronal tissue. Activation of these proteins leads to activation of the MAPK and PI3K pathways (80). Biologic models found that fusions lead to oncogenic addiction irrespective of tissue of origin. These gene fusions are rare, found in an estimated 0.2% to 2.4% of all CRCs, but may be enriched in subpopulations, specifically the MSI-H cohort (3%) (81). The landmark 2018 NEJM paper evaluated the efficacy of larotrectinib, a highly selective small molecule inhibitor of three TRK proteins, in 55 adult and pediatric patients with NTRK fusions. Among all patients, the overall response rate was 75% with 55% remaining progression free at 1 year. Three patients had colon cancer, 2 of whom had an objective response. Following receipt of this data, the FDA granted accelerated approval of larotrectinib for all solid tumors. This is only the second time the FDA has approved a drug, independent of cancer type. The NCCN has since followed suit and now recommends testing for NTRK gene fusions and has incorporated larotrectinib as a possible treatment option in guidelines due to its emergence as a biomarker of predictive response in mCRC (82). While we agree with the recommendations to test, more outcomes data is needed for patients with CRC. Furthermore, considering the extremely low prevalence in CRC, one way to increase the yield of identifying this biomarker is to strategically screen in MSI-H patients.
Emerging molecular biomarkers in CRC
Consensus molecular subtypes (CMS)
As the diverse biology in CRC is increasingly appreciated, so are our efforts to further classify molecular subtypes to inform individualized treatment efforts. With this in mind, The Cancer Genome Atlas (TCGA) evaluated >276 colorectal carcinoma samples using exome sequencing, DNA copy number, promoter methylation, messenger RNA and microRNA expression. Their findings demonstrated that 16% of CRC are hyper mutated, with most of these representing MSI-H primarily with hyper methylation and MLH1 silencing. However, 25% of these patients had somatic MMR gene and POLE mutations. In all other patients, the TCGA found recurrent alterations in the WNT, MAPK, PI3K, TGF-β and p53 pathways. Molecular alterations in the WNT pathway were found in 94% of patients, most prominently APC. Regardless of mutations, nearly 100% of tumors had alterations in MYC transcription and confirmed an important role of MYC in CRC (83).
Building on these efforts, the International Colorectal Cancer Consortium classified patients into 4 defined CMS, CMS-1, CMS-2, CMS-3, and CMS-4 (84). Data was assembled from six CRC subtyping algorithms which were each developed independently using gene expression sets. CMS-1, the MSI immune, represents 14% of all patients and have higher rates of MSI, CpG island methylator phenotype (CIMP) high, hypermutation, and BRAF mutations. While the landscape is changing, these patients have historically had worse survival rates after relapse. CMS-2, or canonical subtype, represents 37% of patients, and they typically have higher rates of somatic copy number alterations and high rates of WNT and MYC activations. CMS-3, the metabolic subtype, represents 13% of all patients. This group represents those with lower rates of copy number alterations, high rates of KRAS mutations and metabolic dysregulation. CMS-4, the mesenchymal subtype, represents 23% of all patients and are those with high rates of TGF-β activation and angiogenesis. CMS-4 class tends to have worse OS (84). An additional 13% of patients did not fit into a specific subtype.
The development of CMS subtyping reflects an international collaboration to characterize mCRC in a more robust format, one that moves beyond isolated mutational status to a transcriptomic based model that accounts for the interplay between various critical molecular pathways (72). Previous work has revealed the emerging prognostic and potential predictive implications of CMS classification. One study reports CMS-4 as a poor prognostic group irrespective of receipt of adjuvant chemotherapy (85). Additional retrospective analyses also highlight CMS-4 as a potential predictive biomarker due to limited efficacy noted with oxaliplatin and EGFRi, irrespective of RAS status (86). Furthermore, a single hospital series applying CMS subtyping to 409 CRCs, stages I–IV, revealed CMS-4 has a 5-year relapse free survival rate of 47% compared with 67% for CMS-1–3 (87). These prognostic findings highlight the need for innovative biomarker driven trial design for the CMS4 subtype.
Poor outcomes in CMS-4 may be a reflection of the high expression of genes associated with T regulatory cells, myeloid-derived suppressor cells, monocyte-derived cells, and TH17 cells in addition to TGF-β activation, all promoting an immune-excluded tumor microenvironment (88). Therefore, understanding this biology will be critical in designing appropriate immunotherapy-based trials for refractory CMS-4, microsatellite stable (MSS) mCRC patients moving forward. To that end, an increasing number of clinical trials that categorize patients and treatments on the basis of their molecular subtype are planned, with one such effort currently underway (NCT03436563).
Despite this prognostic signal, CMS classification is not yet currently included in staging or consensus guidelines with any recommendations based on specific subtypes. However, we expect that CMS subtyping will continue to refine trial design, fostering biologically defined rational drug combinations aimed at patients who are most likely to have a favorable response.
PIK3CA
Phosphatidylinositol-4,5-bisphosphonate 3-kinases (PI3K) are lipid kinases that regulate signaling pathways downstream of EGFR. The PI3K signaling pathway is important in carcinogenesis of multiple cancers. Particularly, PIK3CA (the phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha gene) increases downstream AKT-mTOR signaling pathways which promotes cell proliferation, motility and survival (89). Additionally, upregulation of PI3K increasing prostaglandin E2 synthesis, results in inhibition of apoptosis in colon cancer cells (90). In CRC, PIK3CA mutations are found in 10–20% of tumors and are associated with MSI-H, KRAS mutations and poorer prognosis (91,92). Currently genetic profiling is not routinely recommended; however, it is included in broad genomic sequencing and has gained interest as a potential therapeutic target.
Intriguingly, aspirin use has been associated with improved survival in patients with PIK3CA mutated tumors but not in patients who are PIK3CA WT. A retrospective review of two prospective cohort studies found that aspirin was associated with a significantly longer cancer-specific survival (multivariate HR for cancer-related death, 0.18) (93). However, the data is somewhat mixed, and the NCCN believes that colon cancer survivors may consider taking a 325 mg aspirin, although the reduction in recurrences must be weighed against the increase in gastrointestinal (GI) bleeding and hemorrhagic stroke (10).
There are an emerging number of strategies investigating inhibitors targeting PI3K/mTOR pathway. Our institution has demonstrated that the PI3K pathway is associated with increasing immune infiltration and upregulation of immune checkpoints. Current strategies to combine PI3K inhibition with immunotherapy are underway (94). There are a number of additional studies ongoing; however, targeted therapies directed at this pathway are at this point purely investigational.
Non-TRK fusions (ALK, RET)
RET fusions have been described in various solid tumor including thyroid, non-small cell lung cancers and in <1% of CRCs (95). With impressive responses in the other tumor types, there is enthusiasm for use of multi-targeted TKIs with drugs such as regorafenib and cabozantinib or more targeted agents such as RXDX-105. Early data suggests that patients with RET rearrangements have a worse prognosis compared to RET negative patients with a median OS of 14 vs. 28 months. In that same study, a single patient with MSI-H, and a RET fusion received the novel selective inhibitor RXDX-105 and has had a CR and is progression free at 19 months (96). Clearly more data is required, but preliminary data has been encouraging.
ALK rearrangements were first discovered in anaplastic lymphoma but were later discovered in numerous other cancers—most prominently non-small cell lung cancer. ALK inhibitors including crizotinib and ceritinib have become standard of care for patients with lung cancer harboring ALK fusions with superior efficacy compared to cytotoxic medications (97). While less frequent, ALK rearrangements have been observed in 0.05–2.5% of mCRC patients. However, there is intrigue, similar to NTRK, as their presence is felt to represent oncogenic drivers exclusive of RAS or BRAF mutations (98). At this juncture, use of ALK inhibitors is limited to case reports, with encouraging early reports (97). However, widespread use is not yet warranted. Clinical trials are ongoing evaluating their use (NCT03792568).
BRAF non-V600
Atypical, non-V600 BRAF (aBRAF) mutations are a rare molecular subset of mCRC distinct from BRAFV600E (Class I). Pre-clinical data categorized BRAF into those with intermediate-high kinase activity without RAS dependency (class II) and those with low kinase activity that are RAS dependent (class III) (99,100). Alterations in class II or III are less frequent and account for 2.2% of all patients tested or 21.6% of all BRAF mutations in CRC. In contrast to BRAFV600E, patients with aBRAF are more often MSS, left sided, lower grade, not mutually exclusive from RAS mutations and have decreased rates of peritoneal metastatic disease. In the most robust study to date characterizing aBRAF, these patients have a median OS of 60.7 months in contrast to BRAFV600E OS of only 11.7 months (101). There is emerging pre-clinical data that suggests that patients with class III mutations may be sensitive to EGFRi (100). However, retrospective data has been less encouraging to date, with one study demonstrating no responses to EGFRi, regardless of class, with class II emerging as a negative predictive biomarker and detection of aBRAF in circulating tumor DNA (ctDNA) potentially reflecting a novel mechanism of resistance (102). Currently, there is not enough data to fully support the use of EGFRi for both class II and III aBRAF until more prospective data is available. Innovative trial design with novel agents and rational approaches is an area of active investigation and timely for this rare subset of CRC moving forward.
POLE
POLE mutations are an emerging biomarker that may predict response to immunotherapy. The POLE gene is located in 12q24.33 and encodes the proofreading exonuclease domain of polymerase epsilon (103). Pathogenic somatic POLE mutations occur in an estimated 1.0% of CRCs and are mutually exclusive of patients with dMMR (104). Patients who harbor these mutations tend to have hypermutated tumors that harbor increased neoantigen load that may predict a response to immunotherapy (105). Still, there is limited anecdotal data regarding treatment, and as a result, there are no guidelines to date for patients with these mutations.
Conclusions
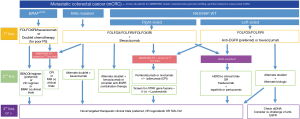
The presence of distinct molecular biomarkers in mCRC influences clinical presentation, histology, guide treatment decisions and therefore directly impact patient outcomes. Identification of established molecular subtypes such as RAS, BRAFV600E, MSI-H, and HER2 amplification in CRC is standard of care, highlights the heterogeneity of this disease and supports the use of precision oncology for refined management (Figure 2). Ongoing studies to unravel and therapeutically target additional biomarkers of clinical and molecular significance such as CMS subtypes, novel amplifications, non-TRK fusions, aBRAF and POLE mutations are paramount to moving the needle forward for this malignancy. Future investigation and novel clinical trial design are necessary to allow for thorough exploitation of emerging molecular biomarkers in terms of identifying their predictive, prognostic and therapeutic potential.
Acknowledgments
None.
Footnote
Conflicts of Interest: The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
References
- Arnold M, Sierra MS, Laversanne M, et al. Global patterns and trends in colorectal cancer incidence and mortality. Gut 2017;66:683-691. [Crossref] [PubMed]
- Bray F, Ferlay J, Soerjomataram I, et al. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 2018;68:394-424. [Crossref] [PubMed]
- Magalhães B, Peleteiro B, Lunet N. Dietary patterns and colorectal cancer. Eur J Cancer Prev 2012;21:15-23. [Crossref] [PubMed]
- American Institute for Cancer Research. World Cancer Research Fund. Diet, Nutrition, Physical Activity and Cancer: a Global Perspective. The Third Expert Report. Available online: http://gco.iarc.fr/today. Accessed April 10, 2019.
- Thun MJ, Linet MS, Cerhan JR, et al. Schottenfeld and Fraumeni Cancer Epidemiology and Prevention. Available online: https://www.scribd.com/document/384238857/Cancer-Epidemiology-and-Prevention-4th-Edition. Accessed April 10, 2019.
- Bailey CE, Hu CY, You YN, et al. Increasing Disparities in the Age-Related Incidences of Colon and Rectal Cancers in the United States, 1975-2010. JAMA Surg 2015;150:17. [Crossref] [PubMed]
- Cronin KA, Lake AJ, Scott S, et al. Annual Report to the Nation on the Status of Cancer, part I: National cancer statistics. Cancer 2018;124:2785-800. [Crossref] [PubMed]
- Souza E, Silva V, Chinen LT, Abdallah EA, et al. Early detection of poor outcome in patients with metastatic colorectal cancer: tumor kinetics evaluated by circulating tumor cells. Onco Targets Ther 2016;9:7503-13. [Crossref] [PubMed]
- Muratore A, Zorzi D, Bouzari H, et al. Asymptomatic Colorectal Cancer with Un-Resectable Liver Metastases: Immediate Colorectal Resection or Up-Front Systemic Chemotherapy? Ann Surg Oncol 2007;14:766-70. [Crossref] [PubMed]
- Kristina Gregory NM, Lisa Gurski O, Benson AB, et al. NCCN Guidelines Version 1. 2019 Colon Cancer Continue NCCN Guidelines Panel Disclosures, 2019. Available online: https://www.nccn.org/professionals/physician_gls/pdf/colon.pdf. Accessed April 11, 2019.
- Van Cutsem E, Cervantes A, Adam R, et al. ESMO consensus guidelines for the management of patients with metastatic colorectal cancer. Ann Oncol 2016;27:1386-422. [Crossref] [PubMed]
- Wolpin BM, Mayer RJ. Systemic Treatment of Colorectal Cancer. Gastroenterology 2008;134:1296-1310.e1. [Crossref] [PubMed]
- Crooke H, Kobayashi M, Mitchell B, et al. Estimating 1- and 5-year relative survival trends in colorectal cancer (CRC) in the United States: 2004 to 2014. J Clin Oncol 2018;36:587-7. [Crossref]
- SEER Survival Statistics - SEER Cancer Query Systems. Available online: https://seer.cancer.gov/canques/survival.html. Accessed April 23, 2019.
- Dhillon AS, Hagan S, Rath O, et al. MAP kinase signalling pathways in cancer. Oncogene 2007;26:3279-90. [Crossref] [PubMed]
- Cox AD, Der CJ. Ras history. Small GTPases 2010;1:2-27. [Crossref] [PubMed]
- Forbes SA, Beare D, Boutselakis H, et al. COSMIC: somatic cancer genetics at high-resolution. Nucleic Acids Res 2017;45:D777-83. [Crossref] [PubMed]
- Cox AD, Fesik SW, Kimmelman AC, et al. Drugging the undruggable RAS: Mission Possible? Nat Rev Drug Discov 2014;13:828-51. [Crossref] [PubMed]
- Piton N, Lonchamp E, Nowak F, et al. Real-Life Distribution of KRAS and NRAS Mutations in Metastatic Colorectal Carcinoma from French Routine Genotyping. Cancer Epidemiol Biomarkers Prev 2015;24:1416-8. [Crossref] [PubMed]
- Simanshu DK, Nissley DV, McCormick F. RAS Proteins and Their Regulators in Human Disease. Cell 2017;170:17-33. [Crossref] [PubMed]
- Carpenter G. Receptors for Epidermal Growth Factor and Other Polypeptide Mitogens. Annu Rev Biochem 1987;56:881-914. [Crossref] [PubMed]
- Saltz LB, Meropol NJ, Loehrer PJ, et al. Phase II trial of cetuximab in patients with refractory colorectal cancer that expresses the epidermal growth factor receptor. J Clin Oncol 2004;22:1201-8. [Crossref] [PubMed]
- Di Marco E, Pierce JH, Fleming TP, et al. Autocrine interaction between TGF alpha and the EGF-receptor: quantitative requirements for induction of the malignant phenotype. Oncogene 1989;4:831-8. [PubMed]
- Jonker DJ, O'Callaghan CJ, Karapetis CS, et al. Cetuximab for the Treatment of Colorectal Cancer. N Engl J Med 2007;357:2040-8. [Crossref] [PubMed]
- Van Cutsem E, Peeters M, Siena S, et al. Open-label phase III trial of panitumumab plus best supportive care compared with best supportive care alone in patients with chemotherapy-refractory metastatic colorectal cancer. J Clin Oncol 2007;25:1658-64. [Crossref] [PubMed]
- De Roock W, Piessevaux H, De Schutter J, et al. KRAS wild-type state predicts survival and is associated to early radiological response in metastatic colorectal cancer treated with cetuximab. Ann Oncol 2008;19:508-15. [Crossref] [PubMed]
- Douillard JY, Siena S, Cassidy J, et al. Randomized, phase III trial of panitumumab with infusional fluorouracil, leucovorin, and oxaliplatin (FOLFOX4) versus FOLFOX4 alone as first-line treatment in patients with previously untreated metastatic colorectal cancer: the PRIME study. J Clin Oncol 2010;28:4697-705. [Crossref] [PubMed]
- Van Cutsem E, Köhne CH, Hitre E, et al. Cetuximab and Chemotherapy as Initial Treatment for Metastatic Colorectal Cancer. N Engl J Med 2009;360:1408-17. [Crossref] [PubMed]
- O'Dwyer PJ, Manola J, Valone FH, et al. Fluorouracil modulation in colorectal cancer: lack of improvement with N -phosphonoacetyl- l -aspartic acid or oral leucovorin or interferon, but enhanced therapeutic index with weekly 24-hour infusion schedule--an Eastern Cooperative Oncology Group/Cancer and Leukemia Group B Study. J Clin Oncol 2001;19:2413-21. [Crossref] [PubMed]
- Loupakis F, Yang D, Yau L, et al. Primary Tumor Location as a Prognostic Factor in Metastatic Colorectal Cancer. J Natl Cancer Inst 2015. [Crossref] [PubMed]
- Tejpar S, Stintzing S, Ciardiello F, et al. Prognostic and Predictive Relevance of Primary Tumor Location in Patients With RAS Wild-Type Metastatic Colorectal Cancer: Retrospective Analyses of the CRYSTAL and FIRE-3 Trials. JAMA Oncol 2017;3:194-201. [Crossref] [PubMed]
- Heinemann V, von Weikersthal LF, Decker T, et al. FOLFIRI plus cetuximab versus FOLFIRI plus bevacizumab as first-line treatment for patients with metastatic colorectal cancer (FIRE-3): a randomised, open-label, phase 3 trial. Lancet Oncol 2014;15:1065-75. [Crossref] [PubMed]
- Douillard JY, Siena S, Cassidy J, et al. Final results from PRIME: randomized phase III study of panitumumab with FOLFOX4 for first-line treatment of metastatic colorectal cancer. Ann Oncol 2014;25:1346-55. [Crossref] [PubMed]
- Schwartzberg LS, Rivera F, Karthaus M, et al. PEAK: a randomized, multicenter phase II study of panitumumab plus modified fluorouracil, leucovorin, and oxaliplatin (mFOLFOX6) or bevacizumab plus mFOLFOX6 in patients with previously untreated, unresectable, wild-type KRAS exon 2 metastatic colorectal cancer. J Clin Oncol 2014;32:2240-7. [Crossref] [PubMed]
- Venook AP, Niedzwiecki D, Innocenti F, et al. Impact of primary (1º) tumor location on overall survival (OS) and progression-free survival (PFS) in patients (pts) with metastatic colorectal cancer (mCRC): Analysis of CALGB/SWOG 80405 (Alliance). J Clin Oncol 2017;34:abstr 3504.
- Brulé SY, Jonker DJ, Karapetis CS, et al. Location of colon cancer (right-sided versus left-sided) as a prognostic factor and a predictor of benefit from cetuximab in NCIC CO.17. Eur J Cancer 2015;51:1405-14. [Crossref] [PubMed]
- Boeckx N, Koukakis R, Op de Beeck K, et al. Effect of Primary Tumor Location on Second- or Later-line Treatment Outcomes in Patients With RAS Wild-type Metastatic Colorectal Cancer and All Treatment Lines in Patients With RAS Mutations in Four Randomized Panitumumab Studies. Clin Colorectal Cancer 2018;17:170-178.e3. [Crossref] [PubMed]
- Davies H, Bignell GR, Cox C, et al. Mutations of the BRAF gene in human cancer. Nature 2002;417:949-54. [Crossref] [PubMed]
- Zhao B, Wang L, Qiu H, et al. Mechanisms of resistance to anti-EGFR therapy in colorectal cancer. Oncotarget 2017;8:3980-4000. [PubMed]
- Cohen R, Cervera P, Svrcek M, et al. BRAF-Mutated Colorectal Cancer: What Is the Optimal Strategy for Treatment? Curr Treat Options Oncol 2017;18:9. [Crossref] [PubMed]
- Jass JR. Classification of colorectal cancer based on correlation of clinical, morphological and molecular features. Histopathology 2007;50:113-30. [Crossref] [PubMed]
- Schirripa M, Lenz HJ. Biomarker in Colorectal Cancer. Cancer J 2016;22:156-64. [Crossref] [PubMed]
- Lai E, Pretta A, Impera V, et al. BRAF-mutant colorectal cancer, a different breed evolving. Expert Rev Mol Diagn 2018;18:499-512. [Crossref] [PubMed]
- Greene C, Atreya CE, McWhirter R, et al. Differential radiographic appearance of BRAF V600E mutant metastatic colorectal cancer (mCRC) in patients matched by primary tumor location. J Clin Oncol 2016;34:abstr 554.
- Seppälä TT, Böhm JP, Friman M, et al. Combination of microsatellite instability and BRAF mutation status for subtyping colorectal cancer. Br J Cancer 2015;112:1966-75. [Crossref] [PubMed]
- Roth AD, Tejpar S, Delorenzi M, et al. Prognostic Role of KRAS and BRAF in Stage II and III Resected Colon Cancer: Results of the Translational Study on the PETACC-3, EORTC 40993, SAKK 60-00 Trial. J Clin Oncol 2010;28:466-74. [Crossref] [PubMed]
- Sinicrope FA, Shi Q, Smyrk TC, et al. Molecular Markers Identify Subtypes of Stage III Colon Cancer Associated With Patient Outcomes. Gastroenterology 2015;148:88-99. [Crossref] [PubMed]
- Stintzing S, Miller-Phillips L, Modest DP, et al. Impact of BRAF and RAS mutations on first-line efficacy of FOLFIRI plus cetuximab versus FOLFIRI plus bevacizumab: analysis of the FIRE-3 (AIO KRK-0306) study. Eur J Cancer 2017;79:50-60. [Crossref] [PubMed]
- Venook AP, Niedzwiecki D, Lenz HJ, et al. Effect of First-Line Chemotherapy Combined With Cetuximab or Bevacizumab on Overall Survival in Patients With KRAS Wild-Type Advanced or Metastatic Colorectal Cancer: A Randomized Clinical Trial. JAMA 2017;317:2392-401. [Crossref] [PubMed]
- Tran B, Kopetz S, Tie J, et al. Impact of BRAF mutation and microsatellite instability on the pattern of metastatic spread and prognosis in metastatic colorectal cancer. Cancer 2011;117:4623-32. [Crossref] [PubMed]
- Phipps AI, Buchanan DD, Makar KW, et al. BRAF Mutation Status and Survival after Colorectal Cancer Diagnosis According to Patient and Tumor Characteristics. Cancer Epidemiol Biomarkers Prev 2012;21:1792-8. [Crossref] [PubMed]
- Fariña-Sarasqueta A, van Lijnschoten G, Moerland E, et al. The BRAF V600E mutation is an independent prognostic factor for survival in stage II and stage III colon cancer patients. Ann Oncol 2010;21:2396-402. [Crossref] [PubMed]
- Van Cutsem E, Köhne CH, Láng I, et al. Cetuximab plus irinotecan, fluorouracil, and leucovorin as first-line treatment for metastatic colorectal cancer: updated analysis of overall survival according to tumor KRAS and BRAF mutation status. J Clin Oncol 2011;29:2011-9. [Crossref] [PubMed]
- Douillard JY, Oliner KS, Siena S, et al. Panitumumab–FOLFOX4 Treatment and RAS Mutations in Colorectal Cancer. N Engl J Med 2013;369:1023-34. [Crossref] [PubMed]
- Maughan TS, Adams RA, Smith CG, et al. Addition of cetuximab to oxaliplatin-based first-line combination chemotherapy for treatment of advanced colorectal cancer: results of the randomised phase 3 MRC COIN trial. Lancet 2011;377:2103-14. [Crossref] [PubMed]
- De Roock W, Claes B, Bernasconi D, et al. Effects of KRAS, BRAF, NRAS, and PIK3CA mutations on the efficacy of cetuximab plus chemotherapy in chemotherapy-refractory metastatic colorectal cancer: a retrospective consortium analysis. Lancet Oncol 2010;11:753-62. [Crossref] [PubMed]
- Seymour MT, Brown SR, Middleton G, et al. Panitumumab and irinotecan versus irinotecan alone for patients with KRAS wild-type, fl uorouracil-resistant advanced colorectal cancer (PICCOLO): a prospectively stratifi ed randomised trial. Lancet Oncol 2013;14:749-59. [Crossref] [PubMed]
- Pietrantonio F, Petrelli F, Coinu A, et al. Predictive role of BRAF mutations in patients with advanced colorectal cancer receiving cetuximab and panitumumab: A meta-analysis. Eur J Cancer 2015;51:587-94. [Crossref] [PubMed]
- Morris V, Overman MJ, Jiang ZQ, et al. Progression-Free Survival Remains Poor Over Sequential Lines of Systemic Therapy in Patients With BRAF-Mutated Colorectal Cancer. Clin Colorectal Cancer 2014;13:164-71. [Crossref] [PubMed]
- Cremolini C, Loupakis F, Antoniotti C, et al. FOLFOXIRI plus bevacizumab versus FOLFIRI plus bevacizumab as first-line treatment of patients with metastatic colorectal cancer: updated overall survival and molecular subgroup analyses of the open-label, phase 3 TRIBE study. Lancet Oncol 2015;16:1306-15. [Crossref] [PubMed]
- Kopetz S, Desai J, Chan E, et al. Phase II Pilot Study of Vemurafenib in Patients With Metastatic BRAF-Mutated Colorectal Cancer. J Clin Oncol 2015;33:4032-8. [Crossref] [PubMed]
- Gomez-Roca CA, Delord J, Robert C, et al. Encorafenib (LGX818), an Oral BRAF Inhibitor, In Patients With BRAF V600E Metastatic Colorectal Cancer (MCRC): Results of Dose Expansion in an Open-Label, Phase 1 study. Ann Oncol 2014;25:iv182-3. [Crossref]
- Corcoran RB, Atreya CE, Falchook GS, et al. Combined BRAF and MEK Inhibition With Dabrafenib and Trametinib in BRAF V600–Mutant Colorectal Cancer. J Clin Oncol 2015;33:4023-31. [Crossref] [PubMed]
- Kopetz S, McDonough SL, Morris VK, et al. Randomized trial of irinotecan and cetuximab with or without vemurafenib in BRAF -mutant metastatic colorectal cancer (SWOG 1406). J Clin Oncol 2017;35:abstr 520.
- Kopetz S, Grothey A, Yaeger R, et al. Updated results of the BEACON CRC safety lead-in: Encorafenib (ENCO) + binimetinib (BINI) + cetuximab (CETUX) for BRAFV600E-mutant metastatic colorectal cancer (mCRC). J Clin Oncol 2019;37:688. [Crossref]
- Kalyan A, Kircher S, Shah H, et al. Updates on immunotherapy for colorectal cancer. J Gastrointest Oncol 2018;9:160-9. [Crossref] [PubMed]
- Le DT, Uram JN, Wang H, et al. PD-1 Blockade in Tumors with Mismatch-Repair Deficiency. N Engl J Med 2015;372:2509-20. [Crossref] [PubMed]
- Jiricny J. The multifaceted mismatch-repair system. Nat Rev Mol Cell Biol 2006;7:335-46. [Crossref] [PubMed]
- Lynch HT, de la Chapelle A. Hereditary Colorectal Cancer. N Engl J Med 2003;348:919-32. [Crossref] [PubMed]
- Halvarsson B, Anderson H, Domanska K, et al. Clinicopathologic Factors Identify Sporadic Mismatch Repair–Defective Colon Cancers. Am J Clin Pathol 2008;129:238-44. [Crossref] [PubMed]
- Koopman M, Kortman GAM, Mekenkamp L, et al. Deficient mismatch repair system in patients with sporadic advanced colorectal cancer. Br J Cancer 2009;100:266-73. [Crossref] [PubMed]
- Venderbosch S, Nagtegaal ID, Maughan TS, et al. Mismatch Repair Status and BRAF Mutation Status in Metastatic Colorectal Cancer Patients: A Pooled Analysis of the CAIRO, CAIRO2, COIN, and FOCUS Studies. Clin Cancer Res 2014;20:5322-30. [Crossref] [PubMed]
- Hutchins G, Southward K, Handley K, et al. Value of Mismatch Repair, KRAS, and BRAF Mutations in Predicting Recurrence and Benefits From Chemotherapy in Colorectal Cancer. J Clin Oncol 2011;29:1261-70. [Crossref] [PubMed]
- Overman MJ, McDermott R, Leach JL, et al. Nivolumab in patients with metastatic DNA mismatch repair-deficient or microsatellite instability-high colorectal cancer (CheckMate 142): an open-label, multicentre, phase 2 study. Lancet Oncol 2017;18:1182-91. [Crossref] [PubMed]
- Overman MJ, Lonardi S, Wong KYM, et al. Durable Clinical Benefit With Nivolumab Plus Ipilimumab in DNA Mismatch Repair-Deficient/Microsatellite Instability-High Metastatic Colorectal Cancer. J Clin Oncol 2018;36:773-9. [Crossref] [PubMed]
- Moasser MM. The oncogene HER2: its signaling and transforming functions and its role in human cancer pathogenesis. Oncogene 2007;26:6469-87. [Crossref] [PubMed]
- Raghav KP, Overman MJ, Yu R, et al. HER2 amplification as a negative predictive biomarker for anti-epidermal growth factor receptor antibody therapy in metastatic colorectal cancer. J Clin Oncol 2016;34:abstr 3517.
- Sartore-Bianchi A, Trusolino L, Martino C, et al. Dual-targeted therapy with trastuzumab and lapatinib in treatment-refractory, KRAS codon 12/13 wild-type, HER2-positive metastatic colorectal cancer (HERACLES): a proof-of-concept, multicentre, open-label, phase 2 trial. Lancet Oncol 2016;17:738-46. [Crossref] [PubMed]
- Meric-Bernstam F, Hurwitz H, Raghav KPS, et al. Pertuzumab plus trastuzumab for HER2-amplified metastatic colorectal cancer (MyPathway): an updated report from a multicentre, open-label, phase 2a, multiple basket study. Lancet Oncol 2019;20:518-30. [Crossref] [PubMed]
- DeStefanis RA, Kratz JD, Emmerich PB, et al. Targeted Therapy in Metastatic Colorectal Cancer: Current Standards and Novel Agents in Review. Curr Colorectal Cancer Rep 2019;15:61-9. [Crossref] [PubMed]
- Pietrantonio F, Di Nicolantonio F, Schrock AB, et al. ALK, ROS1, and NTRK Rearrangements in Metastatic Colorectal Cancer. J Natl Cancer Inst 2017. [Crossref] [PubMed]
- Drilon A, Laetsch TW, Kummar S, et al. Efficacy of Larotrectinib in TRK Fusion–Positive Cancers in Adults and Children. N Engl J Med 2018;378:731-9. [Crossref] [PubMed]
- Cancer Genome Atlas Network. Comprehensive molecular characterization of human colon and rectal cancer. Nature 2012;487:330-7. [Crossref] [PubMed]
- Guinney J, Dienstmann R, Wang X, et al. The consensus molecular subtypes of colorectal cancer. Nat Med 2015;21:1350-6. [Crossref] [PubMed]
- Song N, Pogue-Geile KL, Gavin PG, et al. Clinical Outcome From Oxaliplatin Treatment in Stage II/III Colon Cancer According to Intrinsic Subtypes. JAMA Oncol 2016;2:1162. [Crossref] [PubMed]
- De Sousa E, Melo F, Wang X, Jansen M, et al. Poor-prognosis colon cancer is defined by a molecularly distinct subtype and develops from serrated precursor lesions. Nat Med 2013;19:614-8. [Crossref] [PubMed]
- Sveen A, Bruun J, Eide PW, et al. Colorectal Cancer Consensus Molecular Subtypes Translated to Preclinical Models Uncover Potentially Targetable Cancer Cell Dependencies. Clin Cancer Res 2018;24:794-806. [Crossref] [PubMed]
- Dienstmann R, Vermeulen L, Guinney J, et al. Consensus molecular subtypes and the evolution of precision medicine in colorectal cancer. Nat Rev Cancer 2017;17:268. [Crossref] [PubMed]
- Samuels Y, Wang Z, Bardelli A, et al. High frequency of mutations of the PIK3CA gene in human cancers. Science 2004;304:554. [Crossref] [PubMed]
- Kaur J, Sanyal SN. PI3-kinase/Wnt association mediates COX-2/PGE2 pathway to inhibit apoptosis in early stages of colon carcinogenesis: chemoprevention by diclofenac. Tumor Biol 2010;31:623-31. [Crossref] [PubMed]
- Kato S, Iida S, Higuchi T, et al. PIK3CA mutation is predictive of poor survival in patients with colorectal cancer. Int J Cancer 2007;121:1771-8. [Crossref] [PubMed]
- Benvenuti S, Frattini M, Arena S, et al. PIK3CA cancer mutations display gender and tissue specificity patterns. Hum Mutat 2008;29:284-8. [Crossref] [PubMed]
- Liao X, Lochhead P, Nishihara R, et al. Aspirin Use, Tumor PIK3CA Mutation, and Colorectal-Cancer Survival. N Engl J Med 2012;367:1596-606. [Crossref] [PubMed]
- Nusrat M, Roszik J, Katkhuda R, et al. Association of phosphatidylinositol 3-kinase (PI3K) pathway activation with increased immune checkpoint expression in colorectal cancer (CRC) patients. J Clin Oncol 2018;36:abstr 653.
- Yaeger R, Chatila WK, Lipsyc MD, et al. Clinical Sequencing Defines the Genomic Landscape of Metastatic Colorectal Cancer. Cancer Cell 2018;33:125-136.e3. [Crossref] [PubMed]
- Pietrantonio F, Di Nicolantonio F, Schrock AB, et al. RET fusions in a small subset of advanced colorectal cancers at risk of being neglected. Ann Oncol 2018;29:1394-401. [Crossref] [PubMed]
- Yakirevich E, Resnick MB, Mangray S, et al. Oncogenic ALK Fusion in Rare and Aggressive Subtype of Colorectal Adenocarcinoma as a Potential Therapeutic Target. Clin Cancer Res 2016;22:3831-40. [Crossref] [PubMed]
- Lai AZ, Schrock AB, Erlich RL, et al. Detection of an ALK Fusion in Colorectal Carcinoma by Hybrid Capture-Based Assay of Circulating Tumor DNA. Oncologist 2017;22:774-9. [Crossref] [PubMed]
- Yao Z, Torres NM, Tao A, et al. BRAF Mutants Evade ERK-Dependent Feedback by Different Mechanisms that Determine Their Sensitivity to Pharmacologic Inhibition. Cancer Cell 2015;28:370-83. [Crossref] [PubMed]
- Yao Z, Yaeger R, Rodrik-Outmezguine VS, et al. Tumours with class 3 BRAF mutants are sensitive to the inhibition of activated RAS. Nature 2017;548:234-8. [Crossref] [PubMed]
- Jones JC, Renfro LA, Al-Shamsi HO, et al. Non-V600 BRAF Mutations Define a Clinically Distinct Molecular Subtype of Metastatic Colorectal Cancer. J Clin Oncol 2017;35:2624-30. [Crossref] [PubMed]
- Johnson B, Loree JM, Jacome AA, et al. Atypical, Non-V600 BRAF Mutations as a Potential Mechanism of Resistance to EGFR Inhibition in Metastatic Colorectal Cancer. JCO Precis Oncol 2019;3:1-10.
- Palles C, Cazier JB, Howarth KM, et al. Germline mutations affecting the proofreading domains of POLE and POLD1 predispose to colorectal adenomas and carcinomas. Nat Genet 2013;45:136-44. [Crossref] [PubMed]
- Domingo E, Freeman-Mills L, Rayner E, et al. Somatic POLE proofreading domain mutation, immune response, and prognosis in colorectal cancer: a retrospective, pooled biomarker study. Lancet Gastroenterol Hepatol 2016;1:207-16. [Crossref] [PubMed]
- Giannakis M, Mu XJ, Shukla SA, et al. Genomic Correlates of Immune-Cell Infiltrates in Colorectal Carcinoma. Cell Rep 2016;15:857-65. [Crossref] [PubMed]


